
Koujiro Tohyama
PhD, DVM
Iwate Medical University, Institute for Biomedical Sciences,
Center for Electron Microscopy and Bio-Imaging Research(EMBIR),
Previous Director Laboratory for Nano-Neuroanatomy,
Previous Professor
Analysis of biological specimens by transmission electron microscope (TEM) has entered a new era. Previously, we have to use ultrathin sections with approximate 100 nm-thickness on a grid for TEM observation. In other words, the two different "constraints" of a "grid" and the "thickness of ultrathin section", approximate 100nm, represented limitations for the viable area, and for the precision and depth that could be observed, i.e. not-wide, not-precise, and not-deep, respectively. The ultimate dream of researchers in every field was "wide", "deep" and "precise" analysis of specimens. With recent advances in instrument design, relief from these "constraints" has arrived at last.
Two technologies brought into wide, recent use have provided the solution: Electron beam tomography and backscattered electron imaging. Here we discuss these analytical technologies from our own experience and with some reference to specific data, and consider potential directions for electron microscopy analysis in the life sciences fields.
For more than half a century, ultrathin sectioning has been the starting point for TEM observation. Ultramicrotomes and diamond knives have taken great strides in 30 years, changing ultramicrotomy from a "miracle" to a "common technology". Though a short period of practice is needed even today, this has become an extremely stable technology.
A series of ultrathin sections is obtained on a water-filled diamond knife boat (Fig. 1, left). Their thickness can be determined by interference color (Fig. 1, right). However, the ever-increasing demand for high-resolution analysis in research and examination has brought the following problems to light. Typical TEM projects information for an entire section thickness, showing superposed silhouettes of organelles located at different depth within the section, and 100 nm sections become "too thick" for nano- or subnano- structural analysis. At the same time, as a large series of sections is required to cover an entire structure arranged along the thickness with micrometer scale, these are "too thin" for 3D analysis by reconstruction method. Since the size of ultrathin sections is also typically within 1-2 mm square, specimens are "too small" for research and examination where larger areas of observation are required. Fortunately, methods that overcome these three limitations have now been developed, and some have reached settings accessible even to the ordinary user.

Fig. 1 Series of ultrathin sections obtained with ultramicrotome (left) and chart relating interference color and thickness (right). Sections obtained here are 70-80 nm.
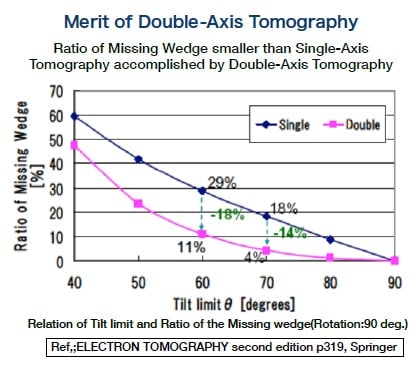
Fig. 2 Difference in missing wedge produced by axis of tilt (single axis and double axis) in electron beam tomography.
This technique is effective for overcoming the "too thick" limitation. A grid holding a section is tilted ±60° with respect to the electron beam, and a series of tilt images is obtained every 1° or 2°. We typically perform double-axis tomography (DA-ET) in which a specimen is rotated 90° within a grid, and the same operation is repeated using two series of tilt images for the same field. In three-dimensional construction, this technique reduces what was roughly a 30% missing wedge area with one axis to an area of approximately 10% with two axes (Fig. 2).
Figure 3 is a three-dimensional reconstruction based on data obtained from DA-ET imaging of a synapse in rat cerebellar cortex. Details are omitted here because these results have not yet been published, but virtually no discrepancies in resolution are seen along the X-, Y-, and Z-axes. Resolution in this instance is 0.58 nm/pixel. Figure 4 shows results from immuno-gold labeling of ultra-cryosection of axon analyzed by DA-ET. The specimen was labeled with ultrafine gold particles (2 nm; used for silver enhanced method) was observed without a silver-enhancement. Immunostaining was provided by antibodies (RT-97) to neurofilament protein 200 kD. The figure shows the same field visualized as a conventional TEM image (left) and a 1 pixel-thick (0.58 nm) image obtained by DA-ET.
In the data for the entire section thickness (left), individual gold particles can be distinguished with some difficulty, but in the image at right, they are clearly discernible. Comparison of the line profile of each set of gold particles (lower figures) makes the difference apparent. Sequential observation of 1 pixel-thick images moving deeper from the surface of the section also shows that neurofilaments alone are labeled specifically at each depth. In general, 5 nm particles at a minimum are used for immunolabeling, and because 2 nm or smaller particles are used for silver enhancement protocol, in this form they are unobserved. This is because distinction from other structures grows more difficult with decreasing size. However, the use of DA-ET allows clear distinction of gold particles, and thus greater resolution in molecular localization. Additionally, we have confirmed that 5 nm gold particles were predominantly localized in the surface of the section, 2 nm and smaller gold particles were found to have penetrated the entire thickness (70 nm in this case) within the ultracryosection (unpublished). The resolution of the labeled image is 0.88 nm per pixel.
Fig. 3 Three-dimensional reconstructed image of a synapse in rat cerebellar cortex analyzed by DA-ET.
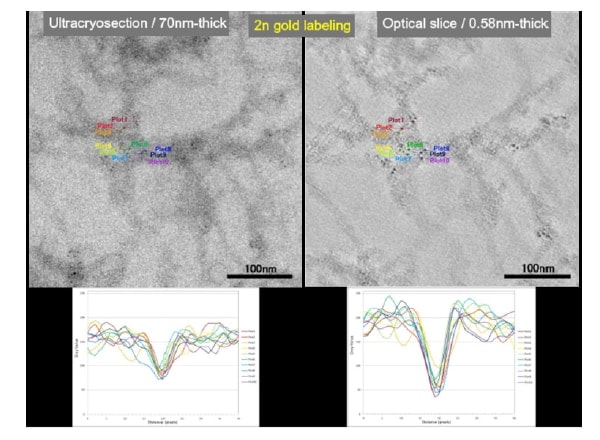
Fig. 4 Line-profile of individual gold particles (color-coded) shown at bottom. Ease of discriminating small-diameter gold particles differs depending on section thickness, and efficacy of information obtained by DA-ET is apparent.
Previously, serial sectioning was the only available means for overcoming the limitation of "too thin" sections. But "axial alignment", aligning the discrepancy in the "axes" of acquired images, was a drawback of serial sectioning. Distortion of individual sections was another major problem. This issue led to the conception and implementation of serial block-face scanning electron microscopy (SB-SEM), a technique in which a specimen surface is sequentially sliced and imaged during a series of image acquisitions, as opposed to imaging of previously sliced sections. A block (specimen)-- in most cases, a resin-embedded specimen-- is placed inside a scanning electron microscope (SEM), and the block face is cut either mechanically or by an ion beam (Ga). Images are acquired primarily by use of back-scattered electron imaging. When specimens are cut mechanically, a miniature microtome is mounted inside the SEM lens tube. A high-vacuum SEM equipped with an ion beam for cutting is known as a dual beam FE-SEM (DB-SEM), and these devices are already in use by manufacturers for quality control of semiconductors and integrated circuits. Figure 5 illustrates the principle of data acquisition by DB-SEM for three-dimensional analysis. Figure 6 presents a three-dimensional image acquired by this method and shows the structure of a rat cerebellar cortex around the Purkinje cell layer. Though the figure shows an assembly of 1-pixel thick images in the X, Y, and Z planes respectively, the information ultimately acquired is volume data and theoretically allows acquisition of a cross-sectional image in any desired plane. However, the original data are not continuance but step images, which limits resolution in the slice orientation to the cutting step (in this case, 50 nm). In this respect, the data differ from tomographic information, which is also volume data. Nonetheless, the thickness of these images is essentially almost half that of conventional TEM sections and provides high resolution. Slices on the order of several nanometers will of course provide highly precise information, but structures of interest must be cut while embedded in the block, making it difficult to determine their location accurately in structures which can only be observed at particular sites. After cutting, a specimen is also expended, and observation and searching cannot be repeated, making this technique unsuited to analysis of rare structures. At present, limited access for ordinary researchers is also a problem.
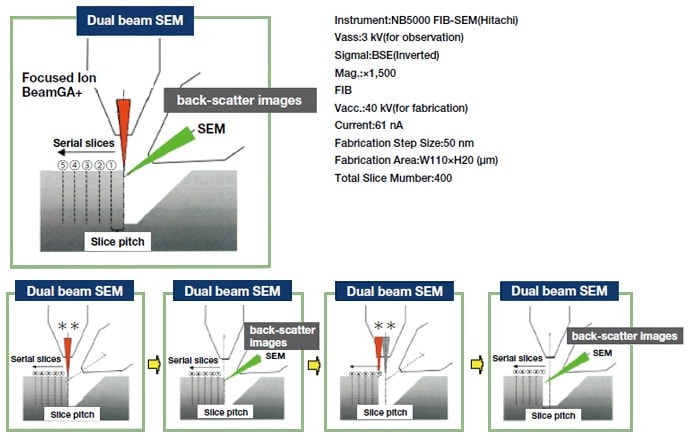
Fig. 5
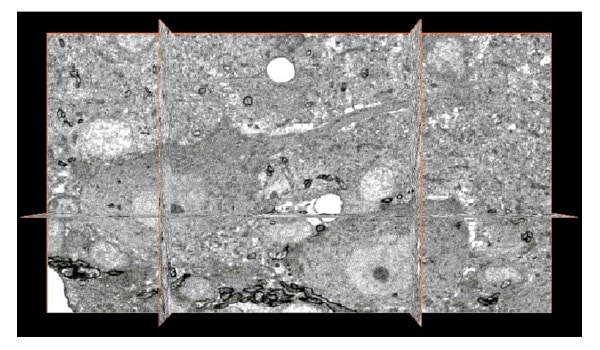
Fig. 6 Three-dimensional reconstructed image of rat cerebellar cortex acquired with HITACHI NB5000 FIB-SEM.
The problem of sections which are "too small" can also be surmounted by the ability to observe large sections. In conventional TEM observation, cut sections are placed on a grid (Fig. 7) for observation, and the observable area is necessarily restricted by the grid. While an one-hole grid allows uninterrupted seamless observation, its area is limited to a diameter of approximately 1 mm (Fig. 7, left). A mesh grid allows observation of a comparatively large area of a cut section (nearly 3 mm diameter), but the image obtained will be discontinuous, since the parts attached to latticework are concealed (Fig. 7, right).
The use of a gridless technique, not employing a grid, would enlarge the area allowing seamless observation. In this technique, ultrathin sections are placed on a slide glass/synthetic resin film or other such carrier sheet and observed by “back-scattered electron imaging” mentioned in the previous section. Figure 8 presents an example of the difference between use of a conventional mesh grid and a gridless technique for serial sections approximately 2 mm wide (4 sections). Without a grid, there are no hidden areas. The width of ultrathin sections which can be cut is limited by the width of the diamond knife used, and the largest commercially available item at present is 8 mm, allowing a section width up to several millimeters. In contrast, the length is limited to the ultramicrotome stroke, which in our experience is a limit of almost 10 mm (Leica EM-UC6). In other words, even the currently allowable size (a several-millimeter square) provides a far more expansive observable area than that in conventional electron microscopy. In practical terms, this technique allows seamless observation of an area approximately 50 times greater than the observable area of approximately 1 square millimeter when using an one-hole grid. Then, areas within a width of several millimeters can be observed by electron microscope without modification, so called "trimming". Now, strict "trimming" is no more necessary, which is another major benefit. Additionally, the same ultrathin section prepared by toluidine blue (TB) staining can also be used for electron microscopy. Usually, we create virtual slides (NanoZoomer, Hamamatsu Photonics) of TB stained section for use in analysis combined with TEM images.
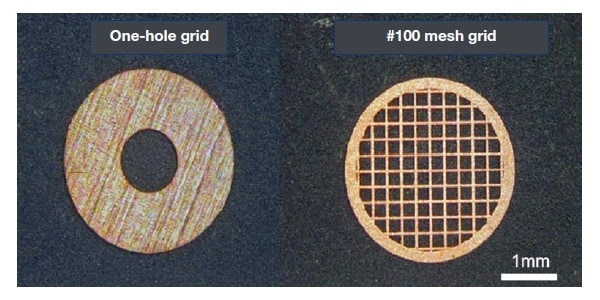

Fig. 7 One-hole grid and mesh grid
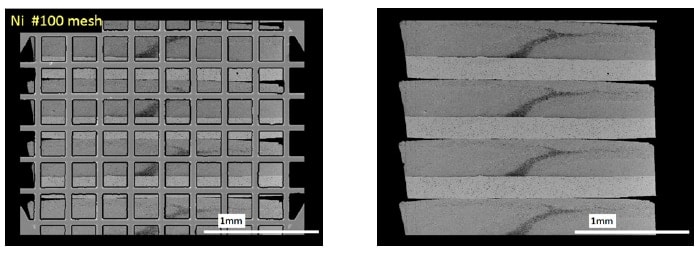

Fig. 8 Comparison of images of four serial sections (2 mm width) on mesh grid (left) and slide glass (right, back-scattered electron imaging). At right, no areas are hidden by latticework of grid, allowing observation of entire section.
A large section obtained was double stained with uranium and lead and then imaged with back-scattered electrons using an SEM (HITACHI SU8010). Imaging parameters were: accelerating voltage 1.5 kV, emission current 20 µA, WD 8.0-2.0 mm, pixel count 2560×1920.
Because back-scattered electron images are typically depicted in positive-negative reverse versus transmission electron images, back-scattered electron images are shown here in reverse (Fig. 9). In the figure, when an image with an expanded section is viewed, any desired site within the specimen bounds can be enlarged. Simple calculation shows that this specimen provides an area for analysis approximately 10 times larger than the area observable with a conventional one-hole grid. Though the resolution of the image does not clearly show the lipid bilayer membrane structure of the cell membrane and is inadequate in other respects as well, it is favorable for basic electron microscopy analysis without a specialized purpose. Notably, digital data obtained can be readily shared with other researchers, which is of great benefit particularly when specimens are valuable. Previously, electron microscopy information even in published papers represented only sites selected subjectively by the authors, and the transparency of the scientific data was not necessarily better than that of data outside the morphology. But a morphological scientific database accessible by researchers worldwide, if created, could increase such transparency substantially in the future, and we recommend use of this technique as one step in the process.
We acquired 10,000-20,000× serial images of wide-area specimens and attempted use of tiling to produce data allowing observation of selected sites within this area up to a point where the original image was enlarged. The image data actually created covered an area of a 2-3 mm square (2.46 nm/pixel), and free image software such as ImageJ (NIH) or NanoZoomer can be used to browse these data. In other words, these data allow electron microscopy observation on an individual PC, even without an electron microscope present. We suggest that this technique be termed "virtual electron microscopy". Use of this technique could, for example, allow use of a sagittal cross-section of the cerebellum to acquire serial data on the Purkinje cell layer alone, over a several-millimeter area, as electron microscopy image data. We have actually used these data to search the microstructure of a murine (mouse) corpus callosum over a comparatively wide area (Reference 3).
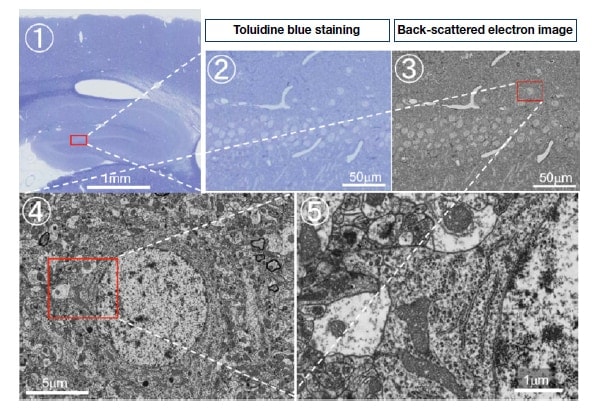
Fig. 9
Example of large-size, ultrathin section observation. Seamless electron microscopy analysis using back-scattered electron image of rat dentate gyrus. Here, these figures demonstrate seamless zooming of a section from mm-scale in Fig.9-(1) up to mm-scale in Fig.9-(5).
Inset figures: Fig.9-(1) shows light microscope image of a part (approx. 3 mm square) of TB-stained large ultrathin section.
Enlargement of red squared area shown in Fig.9-(2).
TB-stained image (Fig.9-(2)) and electron microscope image (Fig.9-(3)) of the same area in the same section.
Enlargement of red square in Fig.9-(3) shown in Fig.9-(4), and enlargement of red square in Fig.9-(4) shown in Fig.9-(5).
Fig.9-(5) exhibits synapses as well as ribosomes, rough endoplasmic reticulum, and other intracellular organelles.
With recent developments in computer technology, axial alignment of serial images has been made comparatively easy. However, it is still difficult to correct image distortion caused by the grids developed for conventional observation of serial sections placed on a grid, and as a result, there is growing use of observational methods in which serial sections are collected on carrier sheets and observed with back-scattered electrons. One group in the very forefront of these efforts is the Lichtman laboratory at Harvard University, which has reported new findings from a complete system these researchers have developed and implemented to analyze enormous numbers of ultramicrotomed, serial sections collected in the form of a ribbon on a special tape (Reference 1). Because the thickness of these sections is approximately one-third that of typical sections used in transmission electron microscopy, they provide high resolution in the Z-axis during reconstruction of serial images. The system also allows collection and analysis of images in thousand-section units.
Depending on the target structure, there may be no need for such a large number of sections, but scarce structures require observation of wide areas. The nodes of Ranvier are one such example. When the nodes are analyzed along the axon, an area of approximately 3-4 microns is collected, requiring 30-40 serial sections.
To analyze central nervous system (CNS) nodes three-dimensionally, we collected 119 sections representing cross-sections of an entire rat optical nerve on a single electroconductively-treated slide glass (Fig. 10) and acquired serial back-scattered electron images including the nodes (40-plus locations). The results showed that astrocyte expected to cover the CNS nodes in textbook fashion accounted for no more than 13% of coverage in this study, and nodes were approximately 76% surrounded by extracellular matrix (ECM) (Fig. 11). Detailed data have not yet been released and are thus omitted here. Though there is growing interest in the importance of the ECM, including the neural nets as well as nodal matrix, a 3-D ultrastructural study of more than 40 nodes has not yet been completed.
As described, our research produced a number of reliable data with relative ease. This work was made possible by the use of SEM back-scattered electrons to image the required number of serial sections in a defined orientation.

Fig.10 Same slide glass holding 119 serial sections. One section 1 mm × 2 mm. Section thickness 100 nm. Relatively well oriented. Also no concerns regarding sequence.
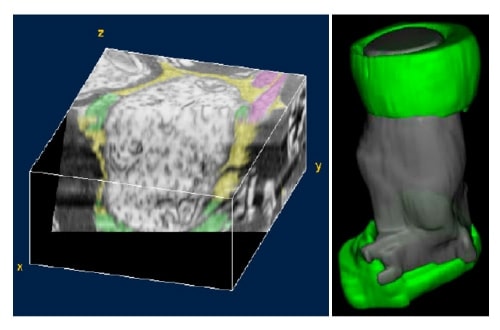
Fig.11 Example of three-dimensional reconstructed image based on back-scattered electron imaging. Green: oligodendrocyte. Pink: astrocyte. Yellow: extracellular matrix and/or space. Left: Cross and longitudinal planes of the node of Ranvier. Surface of the nodal axon is surrounded mainly by extracellular matrix, and in part by astrocyte. Right: Entire surface of nodal axon is surrounded by extracellular matrix without glial elements.
A number of applications using back-scannered electron beam imaging and double-axis tomography are overcoming more than a half century-lasting difficulties and approaching our goal to "finer, wider-area, three-dimensional" analysis in biological sciences. We believe that seamlessly observable, "multi-scale, seamless electron microscopy observation techniques" are no longer a dream, and that the same specimen can be used to obtain data ranging from sub-nanoscale imaging, provided by double-axis electron beam tomography, to scales exceeding several millimeters, provided by ultra-wide area, gridless electron microscopy.
The authors have actually used SB-SEM and DA-ET with pathology specimens to demonstrate special fusion phenomena between plasma cells and dendritic cells (Reference 2).
Additionally, once electron microscope images have been digitized, the constraint whereby electron microscope analysis requires the presence of an electron microscope is increasingly disappearing. The advent of such "virtual electron microscopy" will allow archiving and use of electron microscopy data capable of assuring scientific transparency. This technique likewise opens the gateway to construction of research databases for unprecedented sharing and perpetuation of rare specimens among scientists and researchers. Though we know that even today the silver-impregnated specimens of Santiago Ramon y Cajal, the renowned father of neuroscience, remain preserved in a form allowing electron microscope observation, the advent of this system will provide original specimens that form the foundation for superior research, in turn allowing worldwide sharing of the original electron microscope images. Additionally, in the conceivable instances where reliable experimental techniques are followed to allow wide use of structural images, needless waste of research resources can also be avoided in research using precious experimental animals, without the need for each researcher or research group to secure their own animals and obtain basic data. Moreover, if doubts as to substantive research arise, the capability for widespread access to the raw data will play a part in ensuring transparency of research results and greatly improving trust in scientific research.
Another scenario is also not far-fetched: Ongoing electron microscope data analysis for manuscript revision, with reference to actual electron microscope images. On board the bullet train, destination reached in two hours. Somehow, one-third of electron microscope data needed now complete. Then board an international flight. Take a short break, and in a little more than four hours, acquisition of desired data complete.
Thanks to virtual electron microscopy, there is no longer a need for an actual electron microscope to be present. Electron microscope analysis is possible at any time, in the laboratory office, at home, in transit, and in working accommodations. The potential for such virtual data is now in view. The aggressive development by foreign manufacturers in the field is striking; a 61-beam scanning electron microscope is now a reality, but of course, it is not a technology intended for profit. However, Japanese corporations and technical experts of the past pursued their "dreams"and surprised the world with the unique products they developed. Hopefully, they will build on this experience to energize development of research instruments defining the forefront of analysis in the life sciences and morphology.
Acknowledgements
Research details for this essay were provided with substantial assistance from Kinji Ishida, Chief Engineer, Center for Electron Microscopy and Bio-Imaging Research (EMBIR), Institute for Biomedical Sciences, Iwate Medical University; engineering staff members Tomohito Hanasaka, Eri Matsuura, Masatoshi Ogasawara, and Takayuki Nozaki; and administrative staff member Chikako Asada. Assistance with improvement of SEM specimen stages for slide glass mounting was also provided by Setsuo Maruta, CEO, Nisshin EM Corporation. Through the generosity of Hitachi High-Tech Corporation, we also received the opportunity to use many instruments, active cooperation on technical issues, and improvements to instruments and software. We offer our thanks once again.
Research also included joint research with Professors A. R. Lieberman, W. D. Richardson, P. Anderson, and D. Attwell, University College, London; Professor Yoichi Sugita, Waseda University; Team Leader Yoshio Hirabayashi, RIKEN/BSI; and Eiji Hoshi, Senior Scientist, Tokyo Metropolitan Institute of Medical Science.
Portions of this research were also completed with support from Ministry of Education, Culture, Sports, Science and Technology Grants-in-Aid for Scientific Research (25650181 - Challenging Exploratory Research, 25245069 - Basic A) and the Ministry of Education, Culture, Sports, Science and Technology Support Project for Strategic Research Capability Building at Private Universities.
References
See more