The dugong (Dugong dugon) is a marine mammal in the order Sirenia and family Dugongidae living in shallow tropical seas from the west coast of Africa to the Indo-West Pacific. Japan’s Nansei Islands mark the northern extent of the range of dugongs. Adults grow to about 3 meters in length and weigh up to somewhat less than 500 kg. Dugongs feed exclusively on seagrasses growing in shallow waters (Figure 1). Manatees (Amazonian manatee, West Indian manatee, and West African manatee), another extant member in the order Sirenia, live in the Atlantic Ocean in areas that do not overlap with the range of dugongs. Dugongs are estimated to live up to 73 years. Females give birth after a 14.5-month gestation period. Births are thought to be separated by 3 to 7 years. Newborns nurse for 13.6 to 17.6 months and also consume seagrasses while they nurse1).
Worldwide, the dugong is threatened and listed as vulnerable (VU) on the IUCN Red List of Threatened Species(https://nc.iucnredlist.org/redlist/amazing-species/dugong-dugon/pdfs/original/dugong-dugon.pdf). The Washington Convention on International Trade in Endangered Species limits the trade of derived products. In Japan, dugongs are protected, and hunting restrictions are applied under the national Act on Protection of Cultural Properties, Act on the Protection of Wildlife and Hunting, and Act on the Protection of Fishery Resources as well as the Okinawa Prefecture Ordinance for the Protection of Endangered Wildlife. A recent study found that the East Asian population of dugongs(in the seas around China and Taiwan and Japan’s Nansei Islands), located on the east edge of their range, is at greatest risk of extinction2).

Fig. 1 A dugong swimming (Toba Aquarium)
With extreme hunting practices using dynamite beginning around the turn of the 20th century as documented in fishing records of Okinawa Prefecture and newspaper articles from the time, declines in dugongs in Japan appear to have pushed individual populations below the point of recovery3). Nevertheless, intermittent sightings and bycatch of dugongs near Okinawa Island were reported beginning in the 1970s, and Japanese dugongs appear to have lived in small numbers around Okinawa until at least 2000. Moreover, interviews and submarine surveillance in the Sakishima Islands (the Yaeyama and Miyako Islands) around 2020 indicate that dugongs may still live in those areas. Somehow, the dugong population of the Nansei Islands miraculously and mysteriously survives, possibly helped by additions from migrants from the Philippines and other southern locations traveling in the Kuroshio Current4). Indeed, Okinawan and Philippine dugongs are very genetically close5).
In efforts to protect Japanese dugongs, Japan’s Ministry of the Environment and Okinawa Prefecture have been studying seagrass beds since 2001. (Okinawa Prefecture began doing this in 2016.) Before these studies, experts knew little about the distribution and ecology of Japanese dugongs. Bycatch and stranding information before the studies began suggested that dugongs lived only around Okinawa Island. Since then, however, dugongs have been found to live broadly throughout Okinawa Prefecture, and more information is now available about the seagrass beds that they feed from6). Projects launched to protect dugongs include ongoing efforts to increase awareness among fishermen of techniques for rescuing dugongs trapped in fishing nets to reduce bycatch accidents. Sightings tell researchers more about how the dugong lives, and fishermen have proven to be a valuable source of such information. Okinawa Prefecture has received about 550 reports of sightings to the present (via the Okinawa Dugong Portal Site of the Okinawa Prefecture Environment Department, Nature Conservation Division at https://biodiversity.okinawa/dugong/.) Sightings of dugongs and their feeding trails in Okinawa Prefecture since 2010 are shown in Figure 24).
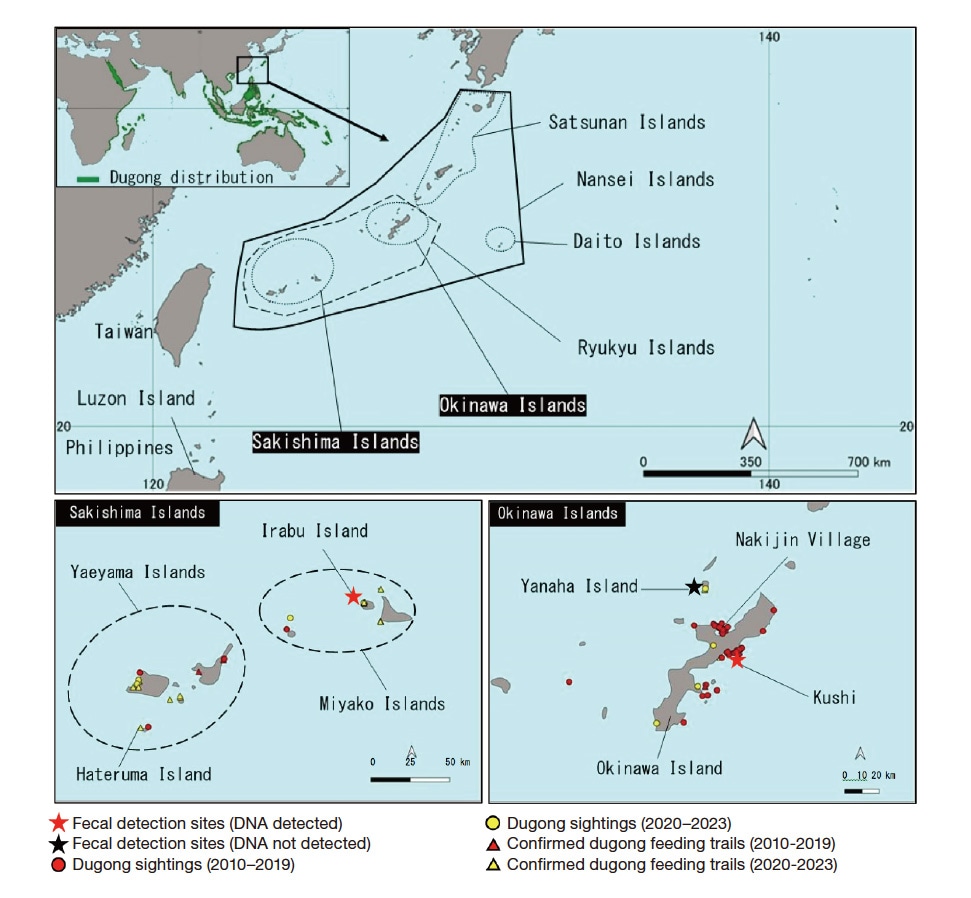
Fig. 2 Locations where feces of large herbivores were collected, animals thought to be dugongs were sighted, and dugong feeding trails were confirmed. The worldwide dugong distribution map at the top left is based on Marsh & Sobtzick, 20197).
To this point, this paper has discussed the basic ecology of dugongs and efforts to protect them in Japan. The distribution of Japanese dugongs and their population sizes remain understudied. In 2022, we began checking for dugong DNA in the feces of large marine herbivores (e.g., dugongs, green sea turtles [Chelonia mydas]) very occasionally found in seagrass beds or washed up on shore. Previous studies had already estimated the distribution of dugongs based primarily on their feeding trails in seagrass beds (Figure 3), but we reasoned that the direct scientific evidence provided by DNA would elucidate the more recent distribution of this animal. Environmental DNA sampling for aquatic animals represents one approach for DNA analysis. This involves collecting water from areas where the target species lives or is raised and checking for DNA in these water samples to understand more about the animal’s life or development. This technique is evolving rapidly in recent years, thanks in part to advances in analytical instruments.
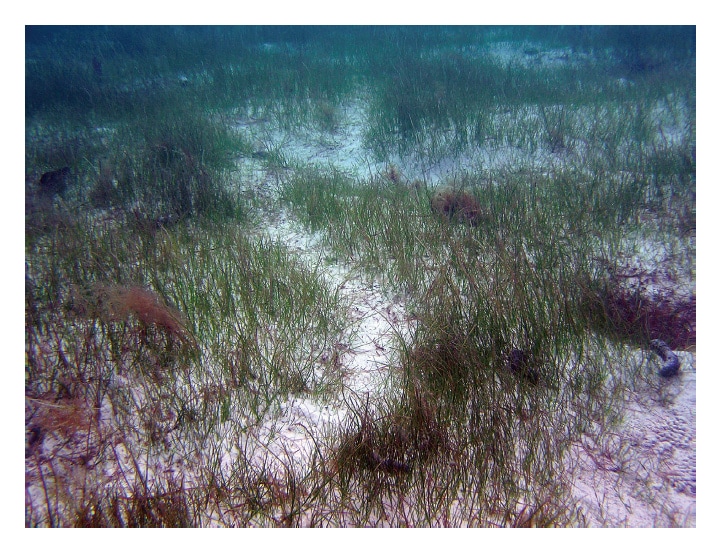

Fig. 3 Feeding trails of dugongs observed in seagrass beds
With the assistance of the Toba Aquarium, we started developing a dugong primer in 2020 capable of specifically amplifying the DNA of this species. Our goal was to detect dugong DNA in seawater in the waters around Okinawa’s islands using an environmental DNA approach8). We used the primer we developed to search for dugong DNA in seawater in the prefecture but, unfortunately, have yet to detect any. We are unsure exactly why but postulate that the extremely small amounts of dugong DNA present in the water mass and the fragile nature of DNA may have contributed. As we were working, a paper was published in 2021 on DNA extracted from the feces of dugongs and green sea turtles in Australia9). Although DNA has been previously sampled from the feces of terrestrial mammals, the approach of the investigators was astonishing in that they used feces of dugongs and green sea turtles collected from the marine environment. Reading the paper, we became convinced that DNA would be easier to detect in fresh feces than in the water mass and decided to use this approach to understand current dugong whereabouts.
Many readers may question why we do not just use aerial surveys to identify dugongs. Indeed, comprehensive and frequent surveys would provide population estimates and other information about Japanese dugongs. Okinawa, however, is surrounded by ocean and contains expansive tracts of seagrass beds along its coastal waters. A comprehensive survey would be prohibitively expensive. Moreover, failing to identify individuals in these minuscule populations would doom a survey, and while dugongs are easy to find in the open ocean, they are very hard to identify in shallow waters with coral sands. Cognizant of these difficulties, we narrow down the area to investigate based on sightings and other information, use drones to image seagrass beds, search for feeding trails and other traces of activity in the images, and then conduct underwater studies based on the findings.
Following is a presentation of a 2024 paper on the identification of dugong DNA in feces collected in Okinawa Prefecture (Irabu Island and Okinawa Island) 4). The study, which was initiated in 2022 and described in this paper, aimed to extract DNA from feces of apparent dugong origin collected in the wild with the goal of identifying new areas where dugongs live.
Feces presumably of dugong origin is sometimes found near seagrass beds and near the surf. Large clumps of feces very occasionally are found on the seabed near seagrass beds, but all feces we found off the shore of Irabu Island in 2022 in which dugong DNA was later detected was floating on the ocean’s surface. The buoyancy of feces likely depends on how long ago it was excreted, what type of seagrass the animal ate, and the gastrointestinal health of the animal. Dugong feces resembles human feces in size and shape, is dark green, and smells remarkably similar to horse and cow feces. Some samples contain small amounts of undigested seagrass fiber (from underground stems and other parts). The feces of green sea turtles, who cohabit areas where dugongs live, is very similar to dugong feces. Green sea turtles also feed on seagrass, and the close similarity of the feces of the two species confounds the efforts of samplers to distinguish them. Not infrequently, large feces apparently from a dugong is revealed by DNA analysis to be from a green sea turtle. We were involved a television special called “In Pursuit of the Elusive Dugong in Okinawa”10), which discussed dugongs off the shores of Irabu Island. The program highlighted the detection of dugong DNA in sampled feces. Many reports of feces sightings and samples have come from many places in the prefecture ever since.
Fecal samples collected on site must be stored in ethanol or another preserving medium and refrigerated as soon as possible to prevent DNA degradation. Ethanol, however, is rarely available to people living on Okinawa’s remote islands to preserve the fecal samples they collect. They instead use the commercially available disinfectant Osvan (10% benzalkonium chloride solution by Alinamin Pharmaceutical), adding 0.5 mL per liter of seawater in which the sample is contained, and then freeze the sample. Now, storage containers, ethanol, and other materials are distributed to islands where fecal samples are often collected. Collected samples are quickly flown frozen to a laboratory on Okinawa Island for DNA extraction soon after arrival. Using the QIAamp Fast DNA Stool Mini Kit (Qiagen) about 100 mg is collected from the surface of the solid part of a fecal sample with a sterile spatula, with about 50 μL of liquid extract obtained. Fecal samples are normally rich in PCR-inhibiting substances, which must be removed with the InhibitEX Buffer in the kit.
We made a dugong-specific primer according to the methodology of the previous Australian study9). The primer sequences were F: 5'-CGCGCGCTATGTACTTCGT-3', R: 5'-GGGGTAAGTAGTGTAATGCACG-3', with a product size of 110 bp. Each sample of extracted DNA was placed in a well and subjected to 2-step PCR. The PCR solution contained 12.5 μL of iProof HF Master Mix (Bio-Rad), 0.5 μL portions of 10 μM primers, 1 μL of DNA solution, and 10.5 μL of sterilized water. The total volume was 25 μL. A Biometra Tone (Analytik Jena) was used as the PCR thermal cycler. PCR was performed by first holding the temperature at 98°C for 45 s and then performing 35 cycles each consisting of 10 s at 98°C, 30 s at 65°C, and 15 s at 72°C followed by an elongation reaction at 72°C for 5 minutes. Ultrapure water was used as a negative control. DNA extracted from the muscle of a dugong carcass found near the coast of the village of Nakijin, Okinawa in March 2019 was used as a positive control. The positive and negative controls were included in each PCR run.
Two-step PCR was performed to enhance the detection of DNA extracted from feces. This involved diluting the first-step PCR products by a factor of 100 with sterilized water and using the resulting solution as a PCR template in the second step. The primers and PCR conditions used in the second PCR step were identical to those used in the first.
The amplification products of the second PCR step were visualized by electrophoresis. A Mupid-exU (Mupid) was used as the electrophoresis device. Agarose gel was prepared by combining 2 g of Agarose S Tablets (Nippon Gene), 6 μL of Midori Green Advance (Nippon Genetics) as a fluorescent DNA stain, and 100 mL of 1× TAE as the electrophoresis buffer liquid. 100 bp DNA Ladder (Takara Bio) was used as DNA molecular weight markers. An Illuminator UltraSlim UV (Gel Company) transilluminator was used to visualize amplification products. To obtain DNA for sequence analysis, products assumed to have a dugong-specific sequence (110 bp) were cut from the agarose gel. The DNA from the gel was processed as directed by the kit instructions using NucleoSpin Gel and PCR Clean-up (Macherey-Nagel). Concentrations of the extracted DNA were determined using the Quantus Fluorometer (Promega) and QuantiFluor dsDNA System (Promega).
Our analyses of feces collected from locations in the prefecture including Irabu Island and Kushi on Okinawa Island produced amplification products apparently of dugong origin (Figure 4) 4).
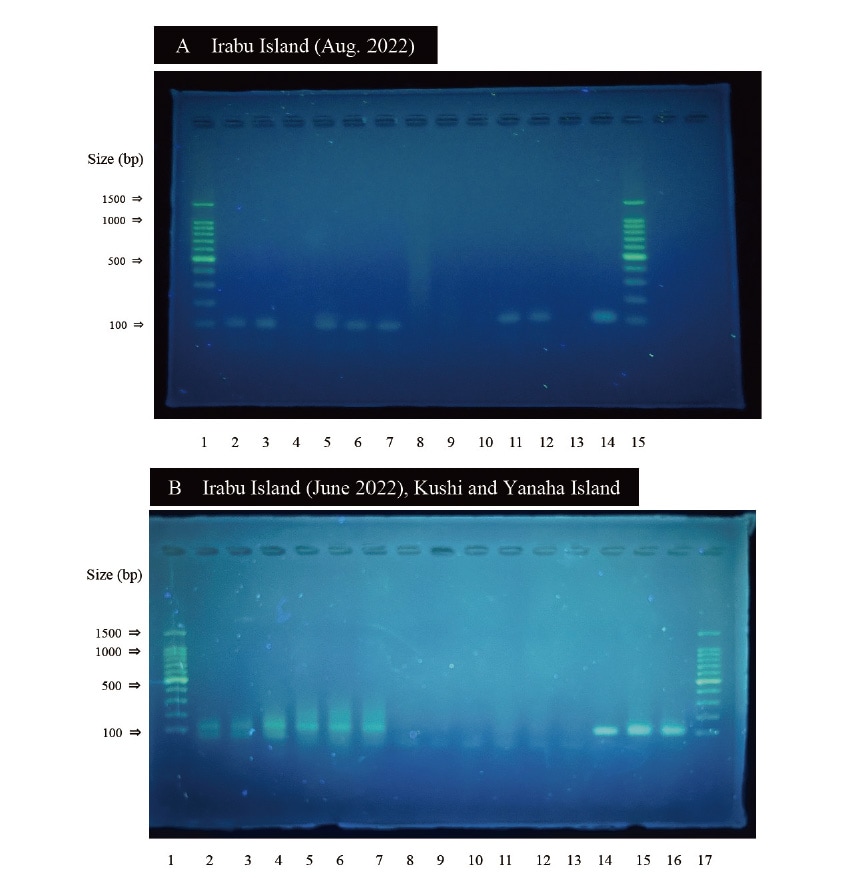
Fig. 4 Photographs of agarose gel electrophoresis patterns of amplicons obtained from PCR of feces using dugongspecific primers. Lanes A1 and A15: DNA ladders, A2-12: from a fecal sample collected from Irabu Island in August 2022, A13: negative control, A14: positive control, B1 and B17: DNA ladders, B2-4: from a fecal sample collected from Irabu Island in June 2022, B5-7: from a fecal sample collected from Kushi in the city of Nago, Okinawa, B8-10: from a fecal sample collected from Yanaha Island, B11-13: negative controls, B14-16: positive controls. Each lane represents a replicate of extracted DNA from a fecal sample. DNA was extracted from the fecal samples shown in Figure 4A as a single replicate and from the samples shown in Figure 4B in replicates of three.
The sequences of the amplification products were determined and then compared against sequences in BLAST (National Center for Biotechnology Information). DNA was fluorescence labeled using the SupreDye v3.1 Cycle Sequencing Kit (EdgeBioSystems). Post-cycle sequence purification was done with the SupreDye XT Purification Kit (EdgeBioSystems). DNA sequence analysis was performed with the DS3000 compact capillary electrophoresis sequencer (Hitachi High-Tech). The sequences determined were compared against dugong base sequence information in BLAST (Figure 5). The sequences of the amplification products completely matched the sequence of dugong DNA registered in BLAST, demonstrating that dugongs still live in the waters off Irabu Island and Kushi on Okinawa Island.

Fig. 5 An operator performs DNA sequence analysis on the DS3000 compact capillary electrophoresis sequencer (Hitachi High-Tech).
The dugong is an umbrella species indicative of biodiversity in tropical seagrass zones. Dugong extinction would lead directly to the degradation of shoreline ecosystems. The fact that a species of this size has managed to survive along the shores of islands in Okinawa is nothing short of a miracle and speaks of the richness of coastal nature there. As we seek to reach the goal of securing a nature-positive world, our generation should shoulder the task of preserving coastal ecosystems, including dugongs and the seagrass beds that nourish them.
Fecal analysis has added scientific evidence of dugong existence, which has until now only been speculated. We will proceed by attempting to use a recently published method for extracting nuclear DNA from feces11) to distinguish individuals and obtain head counts of the still mysterious Japanese dugong subpopulation and also better characterize inter-island migration and other behaviors.
References
See more