
Takashi Fukuda
Ph.D. Agriculture
Associate Professor, Course of Fishery Utilization,
Department of Fisheries, Faculty of Agriculture
Kindai University
The study of pharmaceuticals that use microorganisms as a type of natural resource has made tremendous progress since Alexander Fleming's 1928 discovery of penicillin in Penicillium notatum and the ensuing efforts to make practical use of the discovery. This process spurred the discovery of many pharmaceutical agents that remain in clinical use to this day, including the antibiotics streptomycin and erythromycin, the antifungal agent amphotericin B, the anticancer agents mitomycin and adriamycin, and the immunosuppressant tacrolimus.
Among these pharmaceuticals, the one freshest in the Japanese memory is surely avermectin, for which Dr. Satoshi Ōmura received the 2015 Nobel Prize in Physiology or Medicine1). Avermectin, a secondary metabolite produced by the actinomycete Streptomyces avermitilis, has been used as an anti-parasitic agent that has saved many people at risk for vision loss in Africa. The awarding of the Nobel Prize for this discovery not only celebrated Dr. Ōmura's work but also reconfirmed the importance of research on pharmaceuticals derived from microorganisms.
Meanwhile, the focus of interest in the field of natural resources based pharmaceutical research has recently shifted from terrestrial to marine microorganisms. The number of varieties of microorganisms in the Earth's seas has been estimated to exceed 1 billion, of which more than 90% remain unknown. This suggests that marine microorganisms offer excellent potential for use as materials in pharmaceutical research. For example, clinical trials currently underway in the U.S. are testing the use of pinabulin2) and marizomib3) produced by marine microorganisms, as anticancer drugs. Similarly, seriniquinone4), a compound that exhibits melanoma-selective anticancer activity and was discovered by the author in the culture broth of a marine microorganism has attracted the interest of the American pharmaceutical industry.
As marine microorganisms have recently become popular in drug discovery research, a question that has always vexed natural products researchers is: how can we select–at the earliest possible stages—the strains that produce useful and/or new compounds (that is, strains with the ability to produce novel compounds). Researchers in this area are forever launching new initiatives and studies to answer this question. Examples of attempts to date include studies of bacteria that are difficult to cultivate or symbiotic bacteria, studies of the cultivation conditions required to awaken silent genes, studies that modify genes to promote production of novel compounds, and even methods that obtain genes themselves from the environment to force a variety of expressions in the host microorganisms. However, in all cases these steps must be followed by investigations of the conditions for producing secondary metabolites, and they can hardly be described as shortcuts to the discovery of useful new compounds. Ultimately, the most robust way forward is surely to cultivate microorganisms separated by conventional methods, and then apply a variety of analyses and comparative studies of their secondary metabolites to select those culture broths that give rise to characteristic properties.
Methods for analyzing secondary metabolites in culture broths include comprehensive metabolite analysis using liquid chromatography with UV detectors (LC-UV) or with MS detectors (LC-MS), and molecular-networking methods5) using LC-MS/MS. However, all of these methods require separation of compounds via high-performance liquid chromatography (HPLC) and thus require a certain amount of time to analyze each individual sample. The ideal method would allow culture broths to be analyzed as is with no separation required and would enable visualization (numerical quantification) of the features of individual culture broths. This is our motivation for focusing on fluorescent substances contained in microorganism culture broths.
Fluorescent substances are a class of compounds with the ability to release absorbed light energy in the form of fluorescence. Compounds with this capability, whose structures are known to include benzene or triazole rings, find industrial application in dyes, paints, inks, and research reagents. The detection of compounds in this class proceeds by irradiating a sample with an excitation wavelength and measuring the fluorescence wavelengths emitted in response. For this reason, it may be possible to achieve comprehensive detection of fluorescent substances alone even in substances such as the culture broths of microorganisms containing complicated mixtures of compounds. In this work, we propose a simple method for selecting the culture broths of microorganisms with characteristic features based on the use of fluorescent substances.
The separation of microorganisms used in our experiments was carried out using sediment and marine organisms taken from the East China Sea, the Seto Inland Sea, and the marine environment of the Japanese island of Tanegashima. Supernatant samples (100 µl) diluted by sterilized water were spread onto an agar palate using natural seawater only and cultured at 27°C. After 4 weeks, each colony that appeared on the agar plate was separated and the marine bacteria including actinomycetes (a total of 48 samples) were obtained and stocked.
The cultivation of microorganisms was carried out in two distinct sets of medium: an M1(+) culture medium (using natural seawater) and an M1(-) culture medium (using distilled water). The M1(+) medium consisted of soluble starch (1.0%), Bacto™ yeast extract (0.4%), Bacto™ peptone (0.2%), CaCO3 (0.1%), KBr (0.01%), and Fe2(SO4)3·7H2O (0.004%) dissolved in natural seawater (1 L). The M1(-) medium was identical, but with natural seawater replaced with distilled water. 25 mL of each medium was poured into a 100 mL Erlenmeyer flask, which was then cotton plugged and subjected to sterilization treatment for 15 minutes at 121°C. Then, the media were restored to room temperature and single platinum loops from each of the 48 types of microorganisms (strain numbers 153-200) subject to pure cultivation were inoculated for production culture. Production culture was conducted at 27°C and 160 rpm. After 14 days of cultivation, 25 mL of ether was added and the samples were shaken for one hour to obtain the culture broths (a total of 96 samples).
The culture broths (1 mL) were transferred to an Eppendorf tube and spun for 15 minutes in a 10,000 rpm centrifuge. Then, 300 µl samples of supernatant were transferred to 96 fluorescence-measurement well plates for use as measurement samples.
Measurements were carried out using the instrument of Figure 1 with excitation wavelengths of 200-600 nm to obtain comprehensive measurements of three-dimensional fluorescence spectra at fluorescence wavelengths in the range of 200-600 nm (fluorescence fingerprints).
Instrument used:
F-7100 fluorescence spectrophotometer

Fig. 1 Instrument used for the measurements reported in this paper.
As shown in the experimental data of Figure 2, strong fluorescence was observed in the fluorescence wavelength range of 390-400 nm for the following culture broths: strains 153(-), 153(+), 171(-), 171(+), 188(-), 188(+), 190(-), 191(-), 193(-), and 194(-).
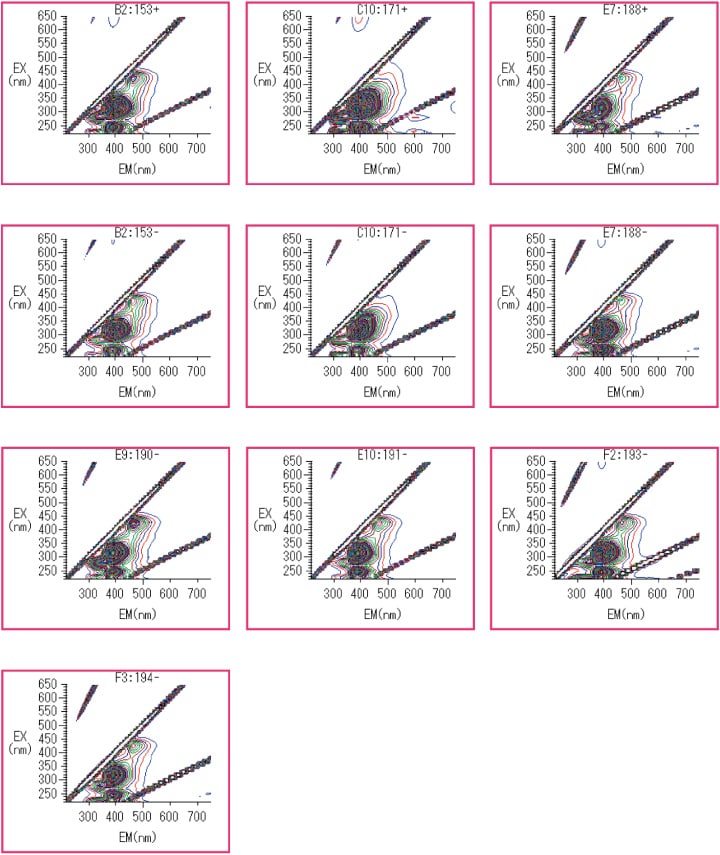
Fig. 2 Culture broths for which strong fluorescence in the range 390-400 nm was observed.
As shown in the experimental data of Figure 3, strong fluorescence was observed at fluorescence wavelengths above 430 nm for the following culture broths: strains 175(+), 176(+), 177(-), 177(+), and 180(-).
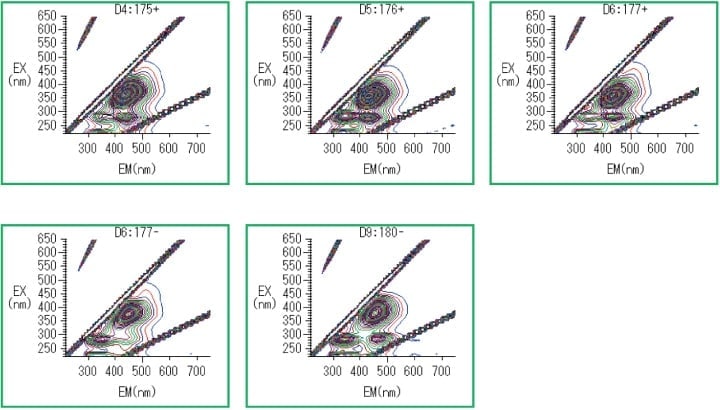
Fig. 3 Culture broths for which strong fluorescence above 430 nm was observed.
Based on the results obtained in Section 3, we select 13 characteristic wavelengths corresponding to peak heights and peak troughs (Figure 4) and use the JMP 12.2 multivariate-analysis software package to conduct a principal-component analysis.
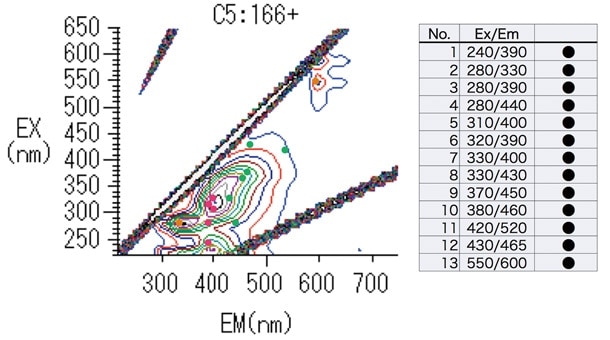
Fig. 4 13 selected wavelengths.
Results are shown in Figure 5. Most culture broths are plotted in the central region of the figure; those that exhibited strong fluorescence in the 390-400 nm wavelength range are plotted in the lower-right portion of the figure (red box), while those for which strong fluorescence above 430 nm were observed are plotted as a group in the upper-right portion of the figure (green box).
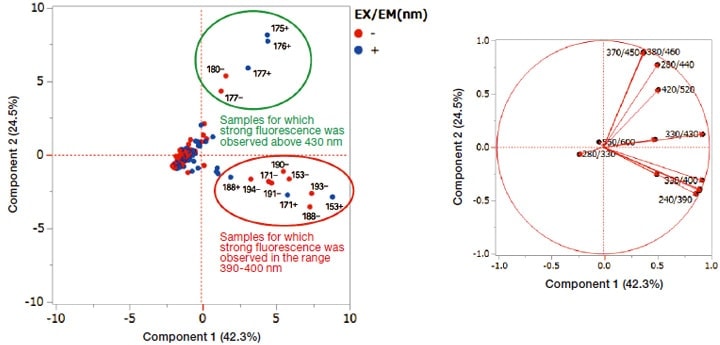
Fig. 5 Results of multivariate analysis.
In this work, we considered the use of multivariate analysis based on fluorescence fingerprints as one method for selecting culture broths containing peculiar marine microorganisms. Using this method, we succeeded in identifying two standout groups of liquids (Figure 5). Strains 171 and 188—regardless of the presence of seawater in the culture broth—were observed to produce compounds with strong fluorescence in the fluorescence wavelength range 390-400 nm; similarly, strain 177 was observed to produce compounds with strong fluorescence above 430 nm, again irrespective of the presence of seawater in the culture broth. These suggest the extremely interesting finding that the respective fluorescent substances may be compounds that mark the corresponding microorganisms as distinctive.
Focusing on the effect of the presence or absence of seawater on the production of fluorescent substances, we see that, among the 10 culture broths with strong fluorescence in the 390-400 nm range, 7 were cases in which seawater was not added. On the other hand, among the 5 culture broths with strong fluorescence above 430 nm, 3 were cases in which seawater was added. Although the small numbers of samples considered in this study prevent us from reaching definitive conclusions, our findings may indicate that culture media containing seawater may promote the production of compounds emitting fluorescence at higher wavelengths.
At present, fluorescence fingerprinting is used in the field of food science for purposes of quality evaluation (to detect contamination by impurities and identify origins of production). The advantages of the method presented in this paper—its ability to make comprehensive measurements of fluorescent substances in samples such as foodstuffs in which various compounds are blended, the extremely simple bare minimum of preprocessing it requires to make measurements, and the enormous quantity of information that it yields, facilitating comprehensive analysis—suggest promising applications to primary screening of foodstuffs (detection of toxic substances)6). In this work, we applied the paradigm of fluorescence fingerprinting to the selection of culture broths of microorganisms and succeeded in rapid selection of strains that produce certain characteristic fluorescent substances. In the future, we expect this technique to be a powerful tool for discovering new and useful substances. Moreover, by combining information yielded by this method with the results of conventional analyses via methods such as LC-UV and LC-MS, we hope to achieve even greater precision in selecting useful strains, a direction we are currently pursuing with collaborators and on which we will report in future publications.
Finally, it is interesting to ask "why do marine microorganisms living in darkness at the bottom of the ocean have the ability to produce fluorescent substances?" We hope that this puzzle will be among the mysteries resolved by future studies of microorganisms.
The research reported here was conducted under the guidance of Prof. Hiroshi Tomoda of the School of Pharmacy at Kitasato University. Marine samples from the East China Sea were provided by Prof. Chiaki Imada of Tokyo University of Marine Science and Technology. In preparing this manuscript, we profited from the cooperation and assistance of Prof. Ryuji Uchida of the Faculty of Pharmaceutical Sciences of Tohoku Medical and Pharmaceutical University. I deeply appreciate all.
References
See more