
Etsuko Utagawa
PhD (Med)
Visiting Researcher, National Institute of Infectious Diseases
Norovirus (Fig. 1) is a small round structured virus (SRSV), which was first detected by Dr. Albert Kapikian of the United States (US) National Institutes of Health (NIH; Bethesda, MD, USA) in patients during a mass outbreak of gastroenteritis at an elementary school in Norwalk, Ohio, USA. Dr. Kapikian used a transmission electron microscope (TEM), which at the time was an exceedingly expensive and specialized piece of equipment, to make this discovery. Blood was taken during the acute and recovery phases from patients who were shedding this SRSV in their stools. Then, using paired serum samples, a definitive diagnosis was made by means of immune electron microscopy (IEM). In those days, genetic testing by polymerase chain reaction (PCR) was not possible, which is a common method used today; thus, investigating causative agents of infectious diseases was extremely difficult. As a result of Dr. Kapikian's discovery, the method of detecting viruses using TEM spread worldwide.
Since before that time, outbreaks of vomiting and diarrheal disease or acute gastroenteritis of non-bacterial origin had frequently been reported in Japan, but the reports attracted little interest outside of Japan because there was no method to detect the causative viruses and the reports were in Japanese. Numerous cases of diseases such as infant cholera, infant vomiting and diarrhea, and infant pseudo-cholera have been reported in Japan since the Meiji Period (1868–1912).
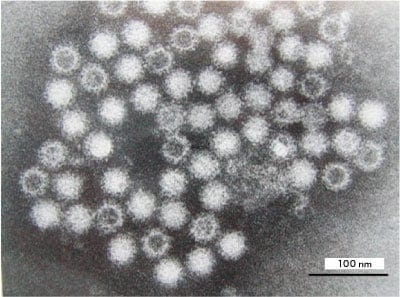
Fig. 1 Transmission electron microscope photograph of human norovirus particles Uranyl acetate staining Bar: 100 nm
Of particular note is a major outbreak of gastroenteritis that affected over 7,000 people in the city of Mobara, Chiba Prefecture, in 1953. This came to be known as Mobara diarrhea1). The cause was a broken sewage basin at a sewage treatment plant that leaked sewage into drinking water at a water supply facility; at the same time, a chlorination malfunction occurred at the supply facility. The municipal government of Mobara City made a request to the then Ministry of Health and Welfare to investigate into the cause of the outbreak, and the investigation was carried out by the National Institute of Health (now the National Institute of Infectious Diseases). Tests by the bacteriology department of the National Institute of Health showed that the diarrhea was not of bacterial origin, but caused by a virus. In order to clarify the incident, medical personnel working on the tests volunteered to take part in a human experiment, in which they ingested fecal material from patients to see whether the same symptoms appeared. Fecal material that had been filter sterilized was demonstrated to cause secondary and tertiary infections, and the report released by the Ministry of Health and Welfare concluded from the reports of chronological monitoring of the infection that the cause of the disease was probably a virus. Unfortunately, however, the various testing methods that were available at that time were unable to actually detect virus particles. Similar records exist in countries all over the world of acute gastroenteritis outbreaks that were believed to be of viral origin.
As mentioned above, in 1969, Dr. Kapikian discovered a virus that measured approximately 27 nm in diameter in fecal material from patients through TEM observation, which at the time was a cutting-edge device, and he published his findings in 1972. In his paper, Dr. Kapikian records that volunteers who took filter-sterilized SRSV became infected, and the same SRSV was found in fecal material from infected individuals by electron microscopy observation. Moreover, he also records that a definitive diagnosis was made using the IEM method with paired serum samples from patients. Based on these results, Dr. Kapikian reported that the SRSV was the cause of the outbreak. Since then, countries around the world, including Japan, have vigorously searched for SRSVs using TEM. As other detection methods have yet to be established, this method has come to be used by institutions such as local public health institutes and universities as the main method of examination when searching for viruses. Many SRSVs have been named after the region where they were detected, such as the Sapporo factor (a virus that caused a mass outbreak of infant gastroenteritis at an orphanage in Sapporo, now known as sapovirus) and the Osaka factor (isolated from patients with gastroenteritis that broke out in Osaka). With more SRSVs being detected by TEM, there were increasing calls for standardization of TEM observation methods. Thus, in 1987, inspection officials from local public health institutes around the country gathered at Tokyo Metropolitan Research Laboratory of Public Health for a range of discussions, which included standardization of patient stool sample preparation for TEM observation, negative staining fluids, and IEM.
As noted above, in 1972, Dr. Kapikian detected SRSV using TEM and showed that this virus was the cause of an epidemic. Reports of SRSV were subsequently made in various countries, but unfortunately, it was still not possible to culture SRSV.
In 1990, the base sequence of the complete SRSV genome was published, which allowed identity screening by means of genetic testing with PCR. Genetic testing by methods such as PCR has become mainstream in recent years. However, because the infectivity of norovirus has still not been clarified, the relationship between gene detection and infectivity is unknown. The meaning of one copy of norovirus genes, which have been reported in numerical terms, remains uncertain, and for this reason the view of the Japanese Ministry of Health, Labour and Welfare is that combined use of TEM and genetic testing is necessary.
Norovirus is an SRSV with a diameter measuring 30–40 nm. In its taxonomic classification, genus Norovirus has only one species, Norwalk virus. However, as the strain of the first ever N. virus to be detected was Norwalk virus, whenever the name "Norwalk virus" is used, it is easy for there to be confusion over whether this means the species or the strain. In addition, the traditional scientific nomenclature for viruses is given in Latin terminations; thus, the virus is currently called Norovirus.
Virus nomenclature is decided by the International Committee on Taxonomy of Viruses (ICTV). According to the ICTV database (ICTVdB), norovirus is defined as a single virus genus belonging to family Caliciviridae. Family Caliciviridae currently comprises five virus genus: Norovirus, Sapovirus, Vesivirus, Lagovirus, and Nebovirus. The virus feline calicivirus that is often used as a substitute for norovirus belongs to family Caliciviridae, genus Vesivirus.
At present, the commonly used term "norovirus" indicates virus attributes and is not a virus species name. If a new virus species belonging to genus Norovirus is discovered, this name will have to be revised. Nevertheless, a Norwalk virus that infects mice is known as "mouse norovirus" and many papers use the term "norovirus" to indicate Norwalk virus. Therefore, the present paper will abide by this custom and use "norovirus."
Norovirus is a non-enveloped virus with single-strand, positive-sense RNA of approximately 7,500 kb, and the genome has three protein coding regions (open reading frames, ORF). ORF1 encodes non-structural proteins involved in replication, ORF2 encodes the viral structural protein VP1, and ORF3 encodes a structural protein. VP1 is a major protein involved in the formation of the virus particle, and P-domain amino acid sequences present on the particle surface have rich diversity and change repeatedly during epidemics. In addition, the norovirus genome is known to recombine at the junction between ORF1 and ORF2. Viruses that have undergone recombination events are known as chimeric viruses.
The norovirus genome sequence has rich diversity and is classified into five genogroups, GI–GV, on the basis of sequence homology. Of these, genogroups GI, GII, and GIV can infect humans.
The majority of viruses identified in cases of human-to-human infection or food poisoning belong to groups GI and GII. Group GIII comprises bovine noroviruses, and the mouse norovirus, which belongs to group GV, was the only virus that could be cell cultured until human norovirus culture was achieved in 2016. In 2015, the original naming system of strains by genotype was revised, which is shown in Table 1 as a reference.
Table 1: Comparative table of norovirus genotypes (from Infectious Agents Surveillance Report (IASR), abridged)
| Old name | New Name |
|---|---|
| GI/1 | GI.1 |
| GI/2 | GI.2 |
| GI/3 | GI.3 |
| GI/4 | GI.4 |
| GI/5 | GI.5 |
| GI/6 | GI.6 |
| GI/7 | GI.7 |
| GI/8 | GI.6 |
| GI/9 | GI.5 |
| GI/10 | GI.8 |
| GI/11 | GI.3 |
| GI/12 | To be determined |
| GI/13 | GI.9 |
| GI/14 | GI.3 |
| New Name | Old name |
|---|---|
| GI.1 | GI/1 |
| GI.2 | GI/2 |
| GI.3 | GI/3 |
| GI/11 | |
| GI/14 | |
| GI.4 | GI/4 |
| GI.5 | GI/5 |
| GI/9 | |
| GI.6 | GI/6 |
| GI/8 | |
| GI.7 | GI/7 |
| GI.8 | GI/10 |
| GI.9 | GI/13 |
| Old name | New Name |
|---|---|
| GII/1 | GII.1 |
| GII/2 | GII.2 |
| GII/3 | GII.3 |
| GII/4 | GII.4 |
| GII/5 | GII.5 |
| GII/6 | GII.6 |
| GII/7 | GII.7 |
| GII/8 | GII.8 |
| GII/9 | GII.9 |
| GII/10 | GII.10 |
| GII/11 | GII.17 |
| GII/12 | GII.12 |
| GII/13 | GII.14 |
| GII/14 | GII.13 |
| GII/15 | GII.16 |
| GII/16 | GII.21 |
| (GII/17=GIV) | – |
| GII/18 | GII.22 |
| GII/19 | GII.15 |
| – | GII.11 |
| – | GII.18 |
| – | GII.19 |
| – | GII.20 |
–:Not applicable.
| New Name | Old name |
|---|---|
| GII.1 | GII/1 |
| GII.2 | GII/2 |
| GII.3 | GII/3 |
| GII.4 | GII/4 |
| GII.5 | GII/5 |
| GII.6 | GII/6 |
| GII.7 | GII/7 |
| GII.8 | GII/8 |
| GII.9 | GII/9 |
| GII.10 | GII/10 |
| GII.11 | – |
| GII.12 | GII/12 |
| GII.13 | GII/14 |
| GII.14 | GII/13 |
| GII.15 | GII/19 |
| GII.16 | GII/15 |
| GII.17 | GII/11 |
| GII.18 | – |
| GII.19 | – |
| GII.20 | – |
| GII.21 | GII/16 |
| GII.22 | GII/18 |
![]()
Genogroup GI is classified into at least 9 species (GI/1–GI/9), and genogroup GII is classified into at least 22 species (GII/1–GII/22). The genotypes fundamentally differ in their antigenicity. Many norovirus genotypes that have been epidemic worldwide in recent years have belonged to GII/4, but there was a major change in norovirus strains that were epidemic in 2015, with GII/4 decreasing in prevalence and GII/17 becoming the main genotype instead. However, GII/4 is presently becoming commonplace again, and there is a tendency for genotypes that become epidemic to vary from year to year.
As norovirus does not have an envelope, it is relatively tolerant to ethanol; thus, alcohol disinfectants are not effective for sterilizing infective norovirus, such as that contained in vomit. Sodium hypochlorite is recommended for sterilizing norovirus. However, it has been shown that heavy use of sodium hypochlorite may negatively impact organisms in the environment (e.g., environmental destruction and pollution), and research to develop an alternative disinfectant is needed.
Treatment of vomitus is essential in cases of food poisoning. Mistakes made in the treatment process when treating vomitus can cause secondary infection, which can easily lead to food poisoning incidents that spread over a wide range. We carried out an experiment to investigate how vomitus falls when someone vomits using simulated vomitus stained with dye and with Bacteriophage Qβ (phage Qβ) added as a substitute for norovirus2). In this experiment, scattering of vomitus was observed by visual inspection of the dye, and virus detection was carried out by assay of viral infection on plastic plates that caught the scatter. Our results showed that simulated vomitus containing dye that fell from a height of 1.6 m scattered concentrically, and scattered material stained with dye was confirmed visually up to a distance of 0.8–0.9 m perpendicularly and 3.1 m horizontally (Fig. 2). The simulated vomitus used in this experiment contained approximately 8 log10 plaque-forming units (PFU)/ml phage Qβ virus. At a distance of 5 m concentrically from the point of vomitus fall, gross inspection of staining did not show any scatter. However, 0.9 log10 PFU/100 cm2 virus was detected on the plastic plates that had been positioned there. These results clarify that there are limits to visual inspection. While no scatter was visually confirmed through observation of dye at 5 m concentrically from the point of vomitus fall, the infection assay showed that vomitus was dispersed over a considerably greater distance than the 2 m distance that had been reported to date. This experiment showed that it is highly likely that norovirus expelled in vomitus is scattered within a radius of 5 m from the point of vomitus fall (the range of scatter was previously considered to be about 2 m). The current descriptions of vomitus treatment by prefectural and regional public health institutes assume virus dispersal within a radius of 2 m, so that the floor and other surfaces between 2 m and 5 m from the point of vomitus fall are not cleaned. There is therefore a very high possibility that dispersed vomitus is present on the floor where sterilization has not been carried out, and that this vomitus is trodden on and spread on shoe soles to places where the vomitus had not been dispersed, thus causing the infectious disease to spread. This is probably a major cause of secondary infection. There are endless cases of infection after treatment of vomitus, and potential dispersal by shoe soles should be borne in mind when treating vomitus. In order to verify this hypothesis, we examined virus dispersal through adherence to shoe soles by using infectivity titer of scrapings from shoe soles and the floor to determine how much simulated vomitus on shoe soles is trodden into the floor as a result of contact between the two3).
In the results of this experiment, which used phage Qβ, once the virus had adhered to shoe soles, it remained stuck to shoe soles up to a walking distance of 49 m. The results thus show that the distance over which the virus is trodden into the floor is approximately 50 m from the point of vomitus fall. The result of this experiment was 50 m, but as the virus remains stuck to shoe soles, there is an undeniable possibility that the virus may be carried even greater distances. This experiment clarified that viruses stuck to shoe soles can be carried a considerable distance. This strongly suggests that infectious viruses present in vomitus that is trodden into the floor are dispersed into the air after drying and become a major cause of secondary infection. Therefore, sterilization over a wide area is imperative to prevent secondary infection. In addition, reports to date have established that inactivation by heat requires heating to a core temperature of 85°C for 1 min or more.
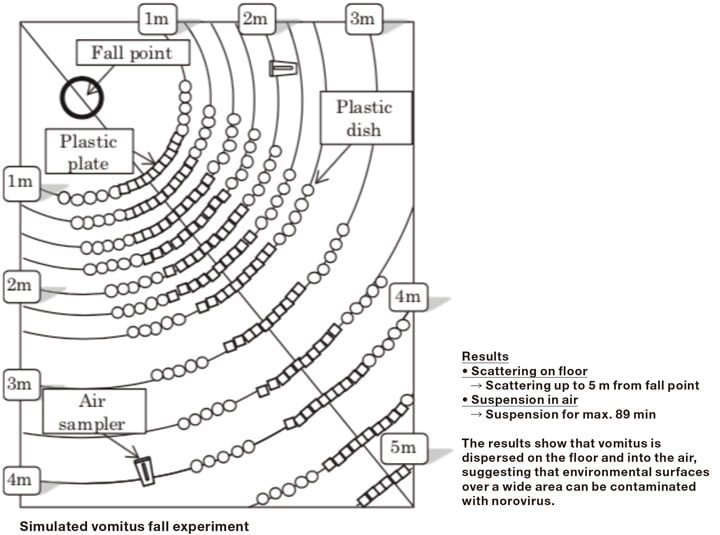
Fig. 2 Evaluation of scattering of simulated vomitus containing phage Qβ
There are a number of points regarding human protective immunity to norovirus that have yet to be elucidated, such as duration or cross-reaction of diverse genotypes, because methods such as neutralization for testing infectiousness have not yet been established. At the individual level, most direct studies involve exposing volunteers to norovirus. The results of such studies show that the duration of immunity to the same virus ranges from about 6 months to 2 years. However, estimates of duration determined by mathematical modeling are longer than these figures.
Characteristic symptoms of norovirus infection include diarrheal disease in adults and vomiting in young children and the elderly. In addition, differences in prognosis are observed between adults and young children and the elderly, who have weaker immune strength. Norovirus infection in young children and the elderly with reduced immune strength is a trigger for aggravation of existing pathologies, and in the worst case can sometimes lead to death. The annual number of deaths from diarrheal disease in undeveloped countries, in particular, exceeds 1 million, and along with other diarrheal disease viruses, norovirus is likely to be a causative virus.
Reports of norovirus epidemic tend to start around the end of October in a normal year and continue until March of the following year. According to the latest Infectious Agents Surveillance Report (IASR) in 2016–2017, reports of norovirus detection began like most years at the start of November and continued to the beginning of March of the next year.
Several advances have been made toward the development of a norovirus vaccine. In the 1990s, Dr. Mary Estes and her group at Baylor College of Medicine (Houston, Texas, USA) successfully manufactured human norovirus virus-like particles (VLP) using an insect cell expression system with a genetically modified baculovirus. Since then, VLP have been used in research into antigenicity of human caliciviruses that cannot be made to proliferate in clonal culture cells, in the manufacture of specific antibodies, in the creation of antigen detection systems, and in research into virus particle morphology. To date, VLP relating to various norovirus strains have been manufactured in different countries. The VLP have the same external structure as human norovirus infective particles and are believed to have the same antigenicity, but no comparison of VLP and infective particles has actually been carried out. However, if analogy with various studies that have been carried out is confirmed, development of a human norovirus vaccine using VLP as vaccine components looks to be a possibility.
From this perspective, Takeda Pharmaceutical Company has joined with the US company LigoCyte Pharmaceuticals to develop a bivalent vaccine that includes GI.1 and GII.4 VLP as a first-generation norovirus vaccine. These vaccines are administered by intramuscular injection and induce antibodies to these types of VLP in the body of vaccinated subjects. In volunteer studies, the induced antibodies have been shown to physically block binding of GI.1 and GII.4 to histo-blood group antigen (HBGA), thus reducing binding efficiency. However, further work is needed to determine whether this virus is effective against genetically different human noroviruses. Many foreign and domestic companies have announced their entry into the field of first-generation vaccines using VLP antigens, and there is considerable hope for research and development aimed at practical application.
In 2016, the Estes group reported in Science that they had succeeded in culturing human norovirus 4). According to their report, human norovirus culture was made possible by using human stem cells. The development of first-generation vaccines using VLP has only just begun, but there is undeniably a possibility that their antigenicities may differ from those of human noroviruses that are actually epidemic in the natural world. It is therefore highly likely that cultivable human norovirus will emerge as an antigen candidate for second-generation vaccines in place of VLP. Nonetheless, countries around the world are competing fiercely to develop a vaccine, and we can be hopeful that possibilities are emerging for the creation in the near future of a vaccine capable of preventing human norovirus epidemics.
References
See more